Upcoming Webinar – Small dyes, big impact: Enabling gentle super-resolution imaging in live cells register now>
PALM microscopy
A Simple Overview
What is PALM microscopy?
PALM is a super-resolution microscopy technique and PALM is short for “Photoactivated Localization Microscopy”. The technique was invented by Eric Betzig along with his colleague Harald Hess and this work was recognized internationally with the award of the 2014 Nobel Prize for Chemistry. Betzig shared the award with Stefan Hell and William Moerner for their contributions to breaking through the diffraction barrier in fluorescence microscopy.
PALM microscopy uses photoactivatable fluorophores to resolve spatial details of tightly packed molecules. Once activated by lasers, fluorophores emit for a short period but eventually bleach. The laser stochastically activates fluorophores until all have emitted. Crucially by activating them randomly and in smaller numbers, a more accurate view of positions can be obtained, making PALM microscopy a key localization microscopy technique.
Signals from each fluorophore are still subject to the 300 nm diffraction of light limit. However, because each has been activated separately, the centre of mass can be calculated accurately. The point spread function (PSF) is used to determine the location down to a resolution of 20 nm. By mapping each of the more accurately defined points together, a complete super-resolution microscopy image can be compiled.
PALM microscopy can be combined with TIRF microscopy or HILO microscopy to improve the signal to noise ratio of thinner sample. This opens up possibilities for researchers studying membrane bound protein complexes or smaller vesicles.
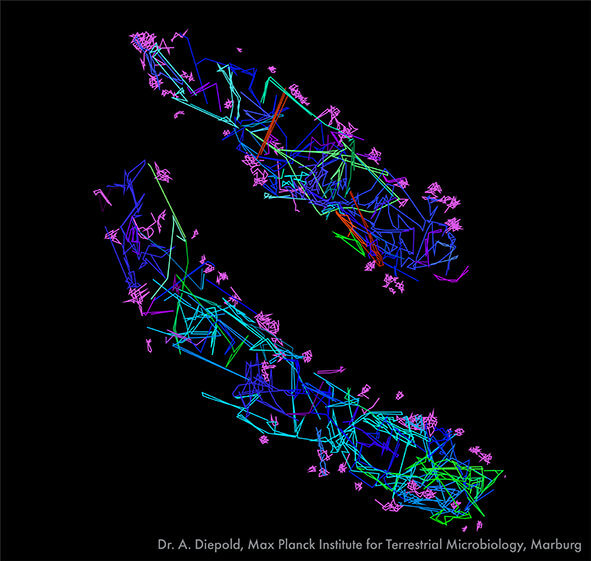
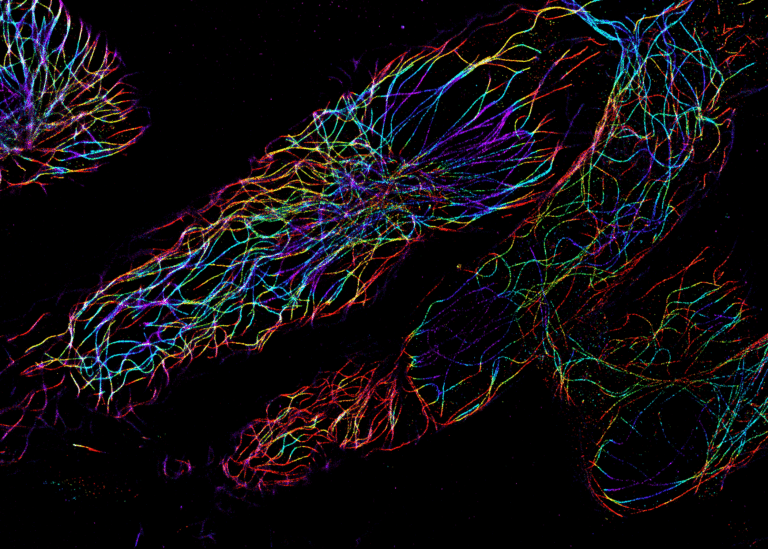
Single-particle tracking of T3SS-HALO stained with JF549 in live bacteria. Sample provided by Dr. A. Diepold, Max Planck Institute for Terrestrial Microbiology, Marburg, Germany. Scale bar is 1µm.
What kind of fluorophores can be used for PALM microscopy?
PALM microscopy and STORM microscopy both use stochastic activation of individual fluorophores to resolve spatial details by identifying precise positions of single molecules. Three commonly used types of fluorophores are:
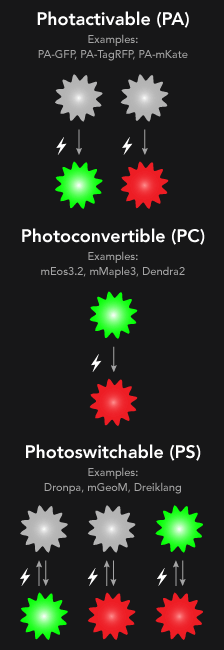
- Photoactivatable Fluorophores: Today most common are photoactivatable proteins, e.g.: PAmCherry, PA-GFP, which emit light upon activation with UV.
- Photoconvertible Fluorophores: Fluorescent proteins which change their emission spectrum upon activation with UV light (e.g.: mEOS proteins).
- Photoswitchable Fluorophores: Typically chemical dyes (e.g. Alexa Fluor 647, DyLight555) which can switch between dark, non-fluorescent and bright, fluorescent states repeatedly.
Read more about the best fluorophores for PALM imaging on our blog. Fluorophores are constantly evolving. New dyes are often photoactivated at higher wavelengths which are more favorable for live samples. Some fluorophores are now available that are both photoswitchable and photoactivatable, providing further options for imaging.

Single-particle tracking of HALO-labeled FtsK protein bound to TMR ligand in Mycobacterium smegmatis cells, samples provided by Zakrzewska-Czerwinska lab, Wroclaw University, Poland. Scale bar is 0.5µm, color scale is 0-6µm2/s.
How can PALM be used?
PALM microscopes can be used for fixed sample localization microscopy to describe the intricate details of analyzed samples with resolutions reaching 20 nm. Using photoactivatable or photoconvertible proteins enables counting of molecules, for example, in complexes and clusters.
Another application is single-particle tracking in live cells. By stochastically activating individual fluorophores we are able to track movement of molecules over time, calculate diffusion coefficients and describe particle behavior.

Single-particle tracking of HALO-labeled FtsK protein bound to TMR ligand in Mycobacterium smegmatis cells, samples provided by Zakrzewska-Czerwinska lab, Wroclaw University, Poland. Scale bar is 0.5µm, color scale is 0-6µm2/s.
Why should the Nanoimager be considered for PALM microscopy?
As a PALM microscope, the Nanoimager‘s extraordinary sensitivity, stability and precision make it an excellent tool for precise measurements, using STORM and PALM microscopy. It is capable of imaging in four colors with two colors running simultaneously, allowing tracking of two distinct fluorophore species at the same time. This allows single molecule localizations and tracks registered in one channel to be relating to cellular markers in the second channel.
The Stage Top Incubator enhances the Nanoimager’s capabilities by providing reliable control over temperature, CO2, and humidity, which are crucial for extending live cell studies. Lasers are digitally controlled providing accurate repeatable results and with a sCMOS camera, images can be generated down to 20 nm resolution. It is a robust, unique design and so can be used in any lab location, not just optical tables in dark rooms. The design naturally dampens vibrations so clean imaging at high resolution becomes possible. The Nanoimager is versatile so it should not be considered as just a PALM microscope. It can also do dSTORM, single-particle tracking and smFRET.
The Nanoimager has been used as a PALM microscope to study RNA polymerase tracking in E. coli, tracking of FtsK in Mycobacterium smegmatis and membrane complexes in bacteria.
Read more about super-resolution microscopy techniques and illumination modes available on the Nanoimager.