Introducing: ONI’s Next-Generation End-to-End Solution for EV Research Learn More
Quantitative Cellular Imaging
Challenge
Quantitatively understanding cells
Understanding how cells work is a primary goal for biological imaging and dSTORM microscopy in particular. For a fundamental grasp on the mechanisms that exist within a cell, a detailed knowledge of how the individual molecules are arranged and how organelles facilitate molecular interactions is absolutely necessary. Furthermore, a quantitative interpretation at this level is needed to precisely measure the effects of external stimuli on the cell, such as the cell’s environment, growth conditions, mutations in the genome or treatment with drugs.
Information that supports this level of interpretation could include the organization of molecules on a membrane, the extent of interaction of different types of molecules, the structural preservation of cellular features and the number and density of molecules in a region.
Solution with the Nanoimager
Using dSTORM microscopy
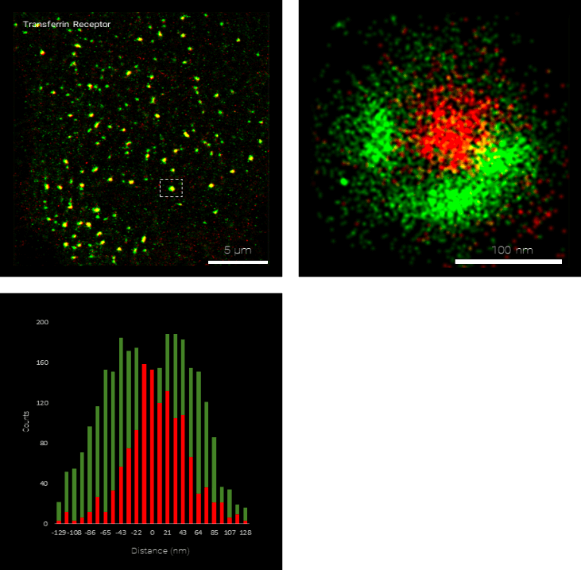
The Nanoimager, and its dSTORM microscopy capabilities, offers the opportunity to see inside the cell or look at purified components with a resolution of under 20 nm using localization-based super-resolution microscopy. This allows quantification and understanding of the internal workings of the cell at an unprecedented level of detail. Taking advantage of the absolute specificity of fluorescence labelling, the spatial relationship of up to four molecular species can be imaged with the 4 laser lines of the Nanoimager. The Nanoimager not only presents super-resolved images of cellular features, but offers several tools for quantifying this information.
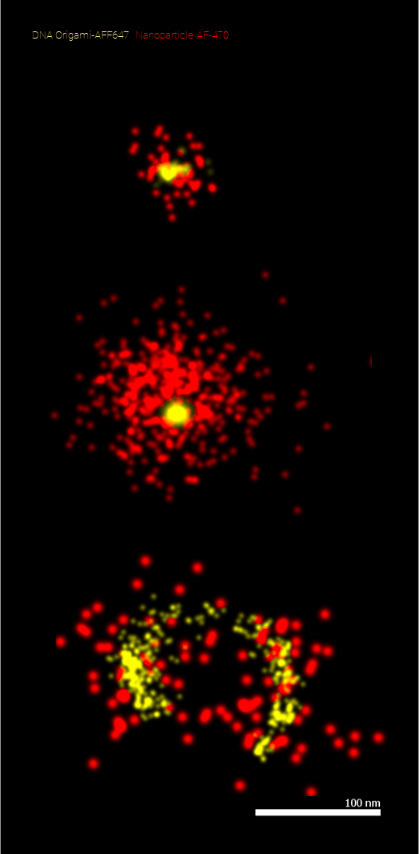
An example application would be imaging the organization of nanomaterials in the therapeutic agents themselves using super-resolution, as shown in the figure.
All of these effects could be in response to a stimulus, a change in the cell environment or a particular stage of the cell cycle. The Nanoimager with it’s extreme sensitivity and easy to use analysis tools can quantitatively discriminate between positive and negative controls. The Nanoimager additionally offers the advantage of a large field of view and simultaneous dual channel imaging, which drastically reduces the time for acquisition and interpretation of results.
Case Study
Analysing mitochondria with dSTORM microscopy
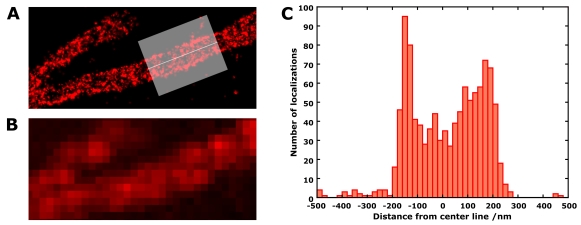
The benefit of super-resolution imaging and dSTORM microscopy is highlighted in this figure, where the shape of mitochondria in neuroblastoma cells was investigated by labeling TOM20 protein. TOM20 was labeled with both AlexaFluor555 and AlexaFluor647 as a control (image at the top). The dSTORM image of a mitochondrion (just the AlexaFluor647 component) is shown in A. The histogram along the axis of the mitochondrion (C) shows the width to be around 300-350 nm, and the denser localizations at the edges caused by the 2D projection of the mitochondrion confirm its tubular structure. This level of information is not obtainable in the conventional widefield image (B).
See more super-resolution microscopy applications and the image analysis and quantification tools available in the NimOS software.