ONIs latest White Paper – Fast, high-throughput LNP sizing and characterization using an automated high-precision workflow read here>

ONI joins the fight against COVID-19
ONI is supporting the global effort into research and diagnostics to tackle the COVID-19 pandemic.
We are currently directing all collaborative and R&D efforts towards this goal.
Our technology is capable of:
- Ultrasensitive, amplification-free detection of proteins and nucleic acids
- Deep phenotyping of virus-cell interactions
- Remote operation, after sample loading
- Full functionality within biosafety cabinets
To discuss your COVID-19 research efforts
Research examples for COVID-19 with ONI technology
How does the virus exploit the host cell machinery?
Ebola hijacks the actin network
These super-resolution images show the effect of Ebola virus-like particles on the actin cytoskeleton. The virus particles co-localize with the disrupted actin network. This reveals how exactly the virus is able to replicate in its host, providing researchers with insights into possible targets for therapeutics.

dSTORM image of actin stained with PhalloidinAF647 and antibodies to nucleoprotein-AF555 in Ebola virus-like particle transfected cells.
A) infected actin
B) healthy cell
C) virus protein nucleoprotein (red) colocalizes with disrupted actin network (blue)
The data was acquired in collaboration with Professor Stephan Becker, a world class virologist at Philipps-Universität, Marburg.
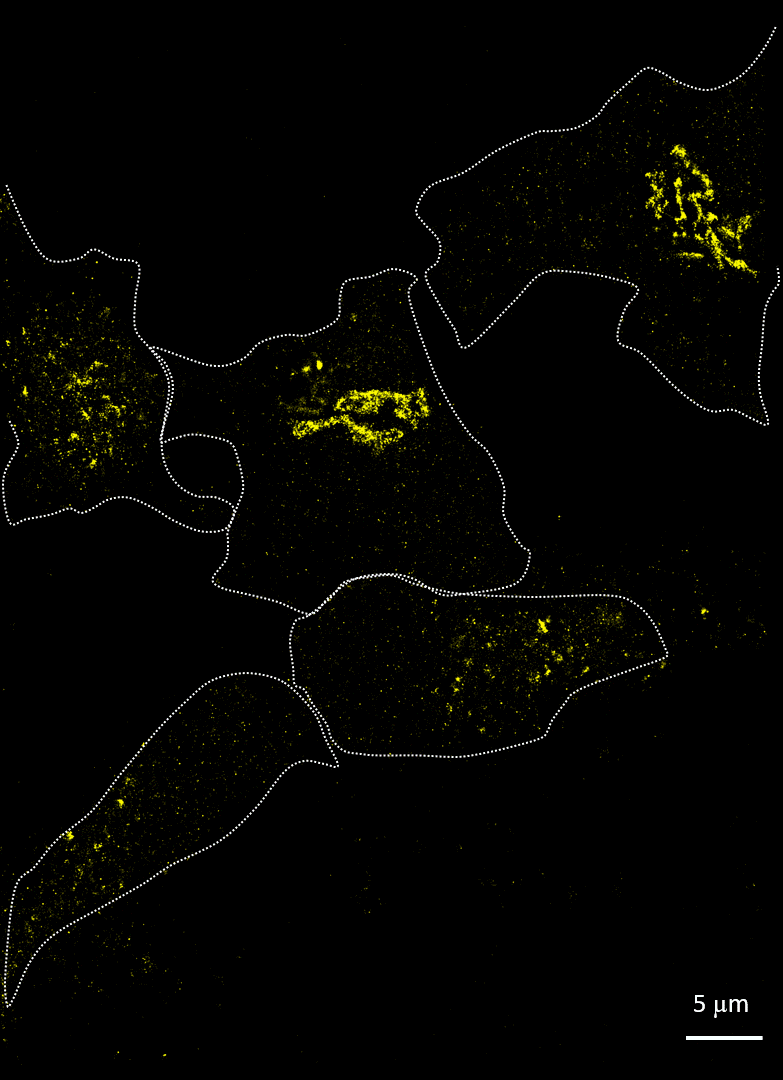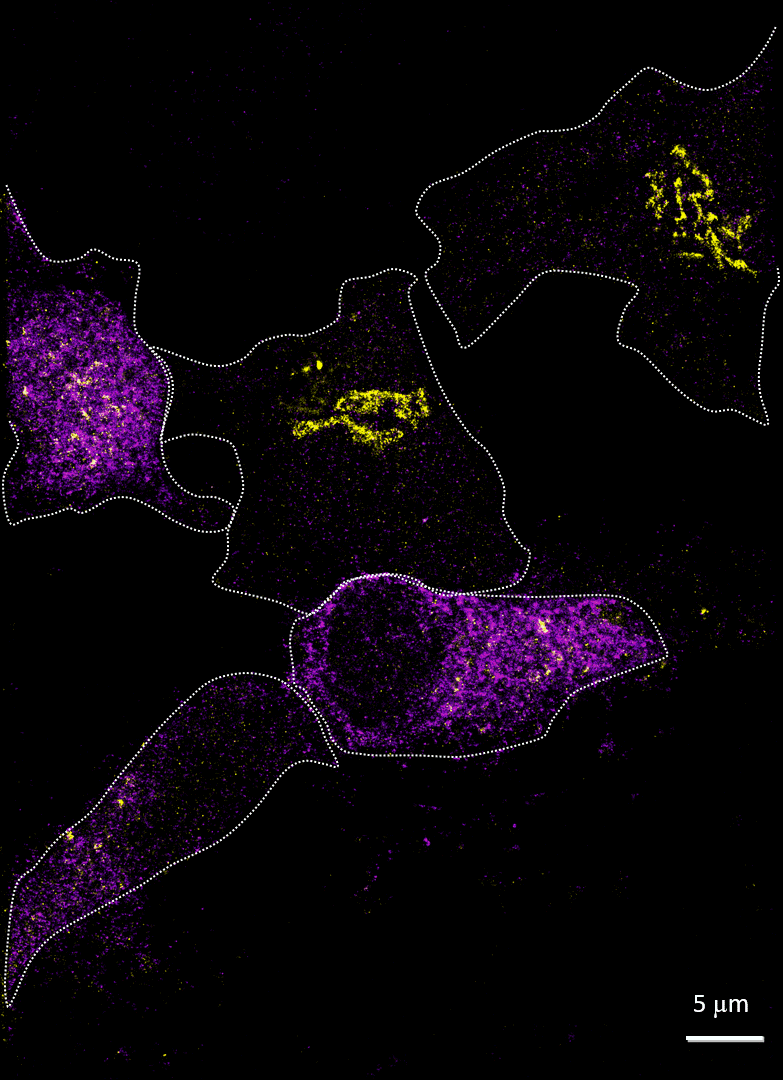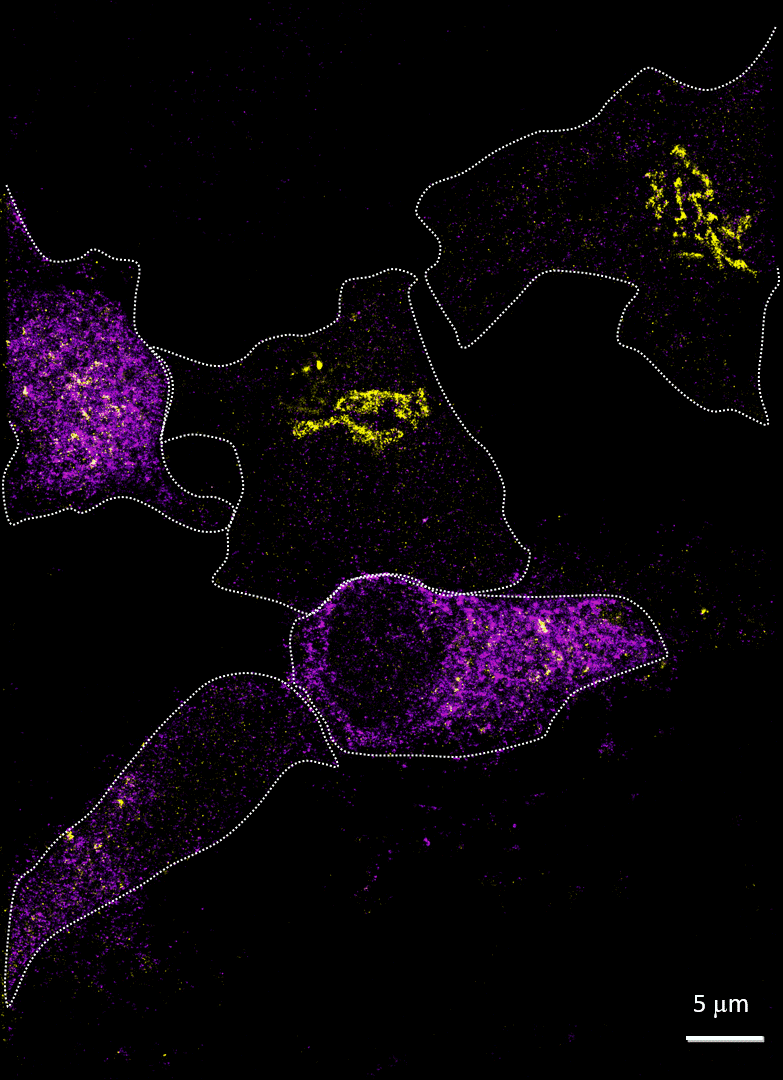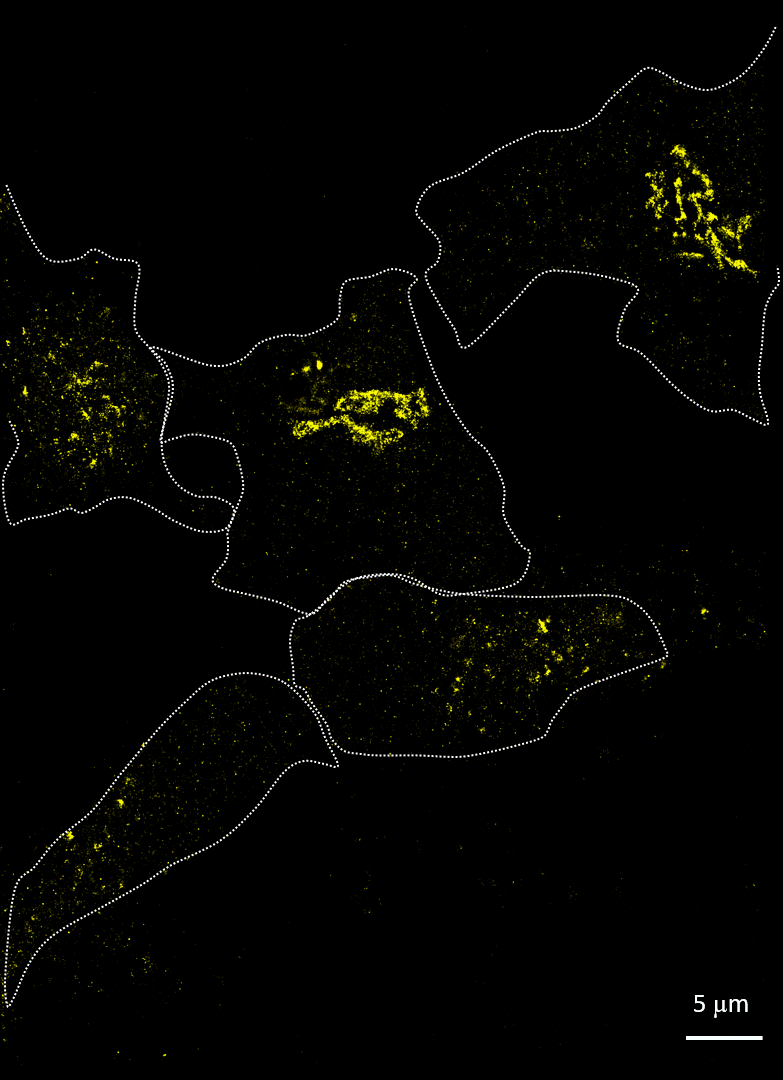
Rhinovirus (viral protein 2C-AF647, purple) infects HeLa cells and disrupts the Golgi (GM130-AF488, yellow). Acquired with Drs A Mousnier and G Schroeder, Queen’s University Belfast
Understanding the effects of viral infection
Rhinovirus disrupts the Golgi apparatus
Imaging cells after infection with a virus can reveal much about the viral mechanism of action. In 2014, Mousnier et al showed how human rhinovirus affects Golgi disruption and protein secretion (doi: 10.1128/JVI.01170-14). We visited Queen’s University Belfast to replicate some of these experiments. Note how the Golgi is left intact in the uninfected cells.

Single-particle tracking of influenza HA (HA-Dendra) inside HEK293T cells, Prof. Salvatore Chiantia, University of Potsdam. Scale bar 5μm
Quantify virus assembly in live cells
Influenza assembles on the plasma membrane
Viruses hijack and exploit their hosts’ cells to replicate and spread. The packaging of new viral particles occurs at the plasma membrane of infected cells. We can understand these assembly sites by visualizing the motion and interactions of key proteins, such as those forming the viral envelope.
In this case we followed single HA proteins, part of the influenza viral envelope, with Professor Salvatore Chianti of University of Potsdam.
Explore
Providing tools to make super-resolution microscopy more accessible
