Upcoming Webinar – Small dyes, big impact: Enabling gentle super-resolution imaging in live cells register now>
7 benefits of using live-cell imaging in single molecule research
The power of live-cell imaging and single-particle tracking
Super-resolution microscopy and particularly Single-Molecule Localization Microscopy (SMLM) are great for capturing molecular events within the nanometer scale resolution. Usually, high resolution is linked to single-molecule imaging of fixed cells or samples. However, additional insights can be obtained from single molecules captured within a live cell imaging context. Here, we review 7 benefits of using live-cell imaging for single-molecule research.
Whether you’re new to super-resolution microscopy or you’re finding your feet in the SMLM journey, you’ve probably asked yourself whether high-resolution live-cell imaging is right for you. In some instances, capturing live events can bring an additional layer of information to super-resolved SMLM images acquired using dSTORM on fixed samples.
For performing SMLM in live imaging the only appropriate technique is Photoactivated Localization Microscopy (PALM), as either dSTORM or DNA-PAINT use solutions that are somewhat toxic to cells. In PALM, photoactivatable or photoconvertible dyes can be used to selectively activate a subset of fluorophores to resolve spatial details of tightly packed molecules. In brief, upon activation with the adequate laser, a sparse number of fluorophores emit for a short period before bleaching. Because fluorophores are only activated at the same time in small numbers until all have emitted, it is possible to then localize and track single molecules over time.
So, why and when should you use super-resolution live-cell imaging? Here are 7 benefits worth considering.
1. Capture dynamic processes at high resolution
As obvious as it may seem, live super-resolution imaging allows you to capture life as the dynamic process it is, at high resolution! This includes investigating molecular processes such as organelle interactions and movements in homeostasis, uptake and release of nanoparticles such as extracellular vesicles, transcription factor dynamics, host-pathogen interactions, cell division mechanisms, or receptor internalization and endocytic transport.
Combined with PALM imaging and Total Internal Reflection Fluorescence (TIRF) microscopy (which only excites fluorophores in the first 200 nm near the surface resulting in improved signal to noise signal), this enables the precise localization and tracking of individual molecules. This provides unprecedented detail on the position and movement over very short time scales, which is critical for understanding rapid biological events.
2. Better understand phenotypes & heterogeneity
Sometimes the ability to study specific phenotypes or capture rare populations that speak of a sample’s heterogeneity is missed with the imaging of fixed specimens. Also, while bulk measurements often mask important differences by providing an average signal from thousands of molecules, single-molecule live-cell imaging allows scientists to see the distribution and variability of behaviors among individual molecules (e.g., differences in diffusion rates or reaction kinetics), which is key to understanding complex cellular pathways.
Some of these phenotypes may occur over time, be highly dynamic, or occur in specific cellular regions, in which case a snapshot might not capture crucial information that could be revealed through live cell imaging.
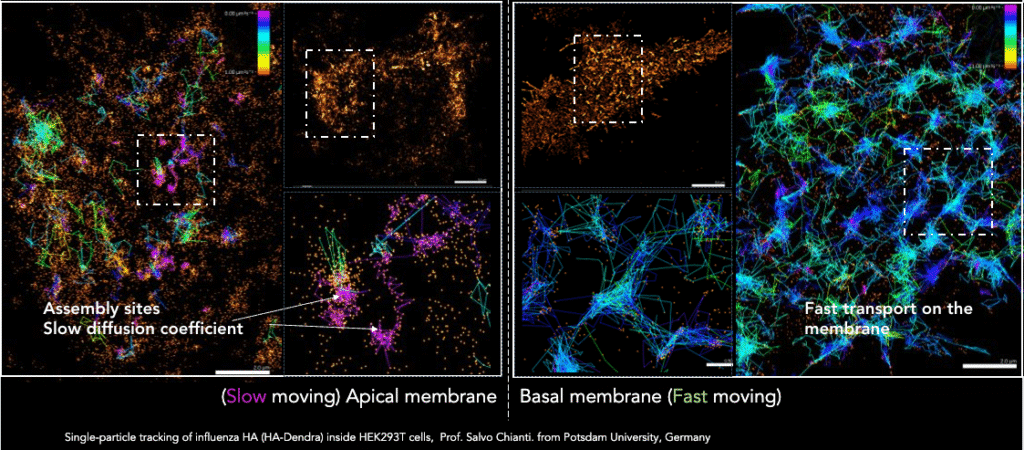
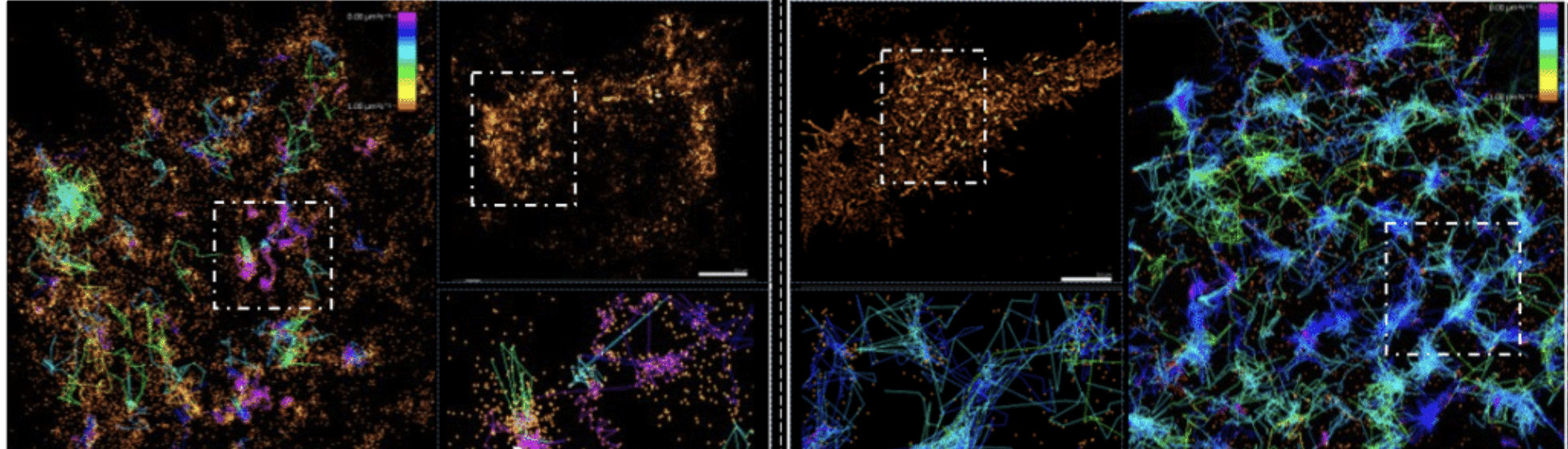
Single-particle tracking of influenza HA-Dendra tagged inside HEK293T cells (provided by Salvo Chianti, Potsdam University, Germany) were at two very different depths: the apical membrane and the basal membrane using different illumination angles. Colored tracks (based on their diffusion-coefficient) show the difference in particle speed depending on their cellular location.
3. Control protein activation as needed
You can gain precise temporal control and turn on a subset of labeled proteins in a specific region of interest or “tag” molecules mid-experiment to follow their movement, using different probes or labeling methods:
- Photoactivatable fluorophores (e.g., PA-GFP): They are variations of commonly used fluorescent proteins that are fused to the protein of interest and emit light upon activation with UV radiation. Note that these are dark by nature and imaging can be tricky at first (the cells will be dark too!), so using a second marker is strongly recommended.
- Photoconvertible fluorophores (e.g., mEos, Dendra): these fluorescent proteins already emit light in their non-converted state. Upon UV light conversion, their emission spectrum changes, offering accurate localization thanks to high photon outputs. Note that the conversion is irreversible.
- Photoswitchable fluorophores (e.g., mGeos): activation is reversible, shuttling between a non-fluorescent and a fluorescent state without photo-bleaching. They tend to be bright, giving high localization precision. Note that to study individual protein dynamics over time in live PALM imaging, it will be better to use photoactivatable or photoconvertible fluorescent proteins that only emit once.
- Hybrid tagging system (e.g., Halo-Tag, SNAP-Tag or CLIP): a combination of genetically-encoded protein tags with a separate cell-permeable synthetic fluorescent ligand. These have gained users in recent years thanks to their efficient and quick labeling, titratable signal using incubation time and concentration of ligand, and strong and durable fluorescent signal.
Want to know which are the best fluorophores for PALM? Read this blog post
4. Track & quantify in real time!
PALM can be harnessed for Single-Particle Tracking (SPT) studies to image and track molecules as they move within cells, recording their motion in a continuous fashion to create a trajectory. This can be quantitatively analyzed for its velocity, diffusion coefficient, displacement, distance traveled, and mean speed. For example, SPT can be used to:
- Visualize interactions in real time
- Measure cargo delivery to organelles
- Assess new methods for drug discovery and delivery
- Elucidate protein docking to specific targets
- And much more!
Together with the use of integrated tracking tools for SPT data analysis, you can easily determine molecular turnover by tracking the activated pool as it degrades or is replaced by new, non-activated proteins, or monitor long-range transport from a known starting point to measure speed and destination with high confidence.
For successful single particle imaging and tracking, it is important to consider and optimize for the density of labeled particles in the sample, the size of the fluorescent tag, the fluorophore brightness, and different labeling methods.
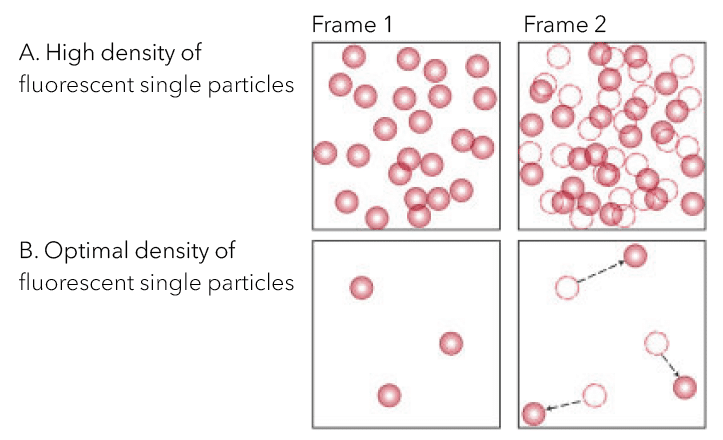
The concentration of labeled particles is critical for trajectory accuracy. (A) High density causes inaccurate tracks due to overcrowding and erroneous linking between frames. (B) Optimal density requires particles to be sufficiently spaced for accurate recording of each particle’s position without neighboring interference.
To find out more about how to get started with Single Particle Tracking: Download our Technical note
5. Study cellular responses to drugs and perturbations
Both PALM and SPT are powerful techniques used to observe and quantify antibody/drug uptake, distribution, delivery, and release within cells. For instance, by tracking individual molecules, you can understand receptor interactions at the nanoscale, aiding in better design and validation of bispecific drugs. The quick imaging capabilities of systems like Aplo Scope or the Nanoimager allow for the capture of target dynamics and the subsequent extraction of properties like diffusion, interactions, and binding kinetics, which are essential for validating drug effect, potency, and cellular response.
The integration of PALM and SPT into automated super-resolution imaging workflows allows for the systematic exploration of drugs or families of proteins, whereby researchers can plate fluorescently labeled cells for imaging, choose a perturbation from a library and apply it with microfluidics, perform live cell imaging, monitor cellular dynamics, and analyze data for desired phenotype then repeat the process!
6. Avoid artefacts induced by fixation and staining
A necessary step in many traditional microscopy methods, chemical fixation, can irreversibly alter cellular structures and the behavior of molecules from their native form, changing their shape and morphology due to osmotic pressures, and leading to potential artifacts. Live-cell imaging tends to bypass this issue and data reflects the molecules’ physiological state within healthy, functioning cells. However, protein tagging can also affect molecular behaviour which needs to be carefully optimized, similarly to the levels of expression (whether it is via transfection or stable transgene expression), as overexpression or dominant negative effects could be observed.
7. Experiment in real time and keep your cells alive
You can also try new experiments, investigate the behavior of molecules under controlled, changing conditions, and have fun seeing the effect in real time… ideally while keeping cells alive. For example, a single experiment can involve adding a ligand to observe receptor-ligand kinetics, or titrating a drug while simultaneously tracking its effects on protein mobility (e.g., using a Halo-Tag label). Live-cell imaging also allows you to use pH-sensitive dyes to observe localized pH changes (i.e., such as those accompanying EV release), or calcium sensors to monitor real-time signaling cascades. These functional probes can be further combined with organelle-specific dyes (like Mitotracker or Cell Mask) to contextualize molecular activity within cellular compartments.
Another key benefit is the ability to optimize signal density on the fly, by adjusting the amount of UV light used for activation in photoactivatable systems, ensuring individual molecules are sufficiently sparse for tracking. Finally, cell preservation after single-molecule live imaging, makes samples available for downstream experiments like Western blot, sequencing, or further FACS analysis, offering a direct correlation between single-molecule dynamics and bulk cellular outcomes.
Having listed all these benefits, there’s something to add…
The biggest challenge in single-molecule imaging lies in navigating the trade-off triangle between spatial resolution, temporal resolution, and sample health. Achieving higher spatial resolution often demands intense laser power and long acquisition times to collect enough photons for precise localization, exemplified by dSTORM. However, this high laser power and extended exposure negatively impacts cell health and imaging time, decreasing temporal resolution and forcing a compromise on the Field of View (FOV) size and frame rate to keep the cells alive. Conversely, rapid live-cell imaging prioritizes temporal resolution at the expense of nanometer-scale detail.
Therefore, fixed-cell super-resolution techniques, like dSTORM, often remain the best option for ultrastructural detail, but the most comprehensive biological answers emerge by treating these methods as complementary tools. For instance, an integrated system like Aplo Scope allows researchers to swiftly acquire real-time dynamics in live cells, then immediately switch to fixed-cell super-resolution imaging to resolve the underlying nanostructure, directly linking molecular function to morphology. The addition of the stage top incubator allows extended live-cell imaging studies with both Aplo Scope and Nanoimager, providing control over temperature, CO2, and humidity to keep live samples in optimal conditions.
Are you ready to start with single-molecule live-cell imaging?
We recommend you watch our recent webinar covering key considerations around sample preparation (e.g., labeling density and homogeneity, cell culture media), image acquisition (e.g., optical setup, illumination angle, frame rate, avoiding phototoxicity), and data analysis (e.g., real time localizer, integrated analysis tools, tracking parameters, exclusion radius, available software solutions, and automated analysis workflows).
If you want to discuss how best to tackle imaging of your targets of interest or any specific steps in the super-resolution imaging workflow, get in touch with our team of scientists at: oni.bio/contact